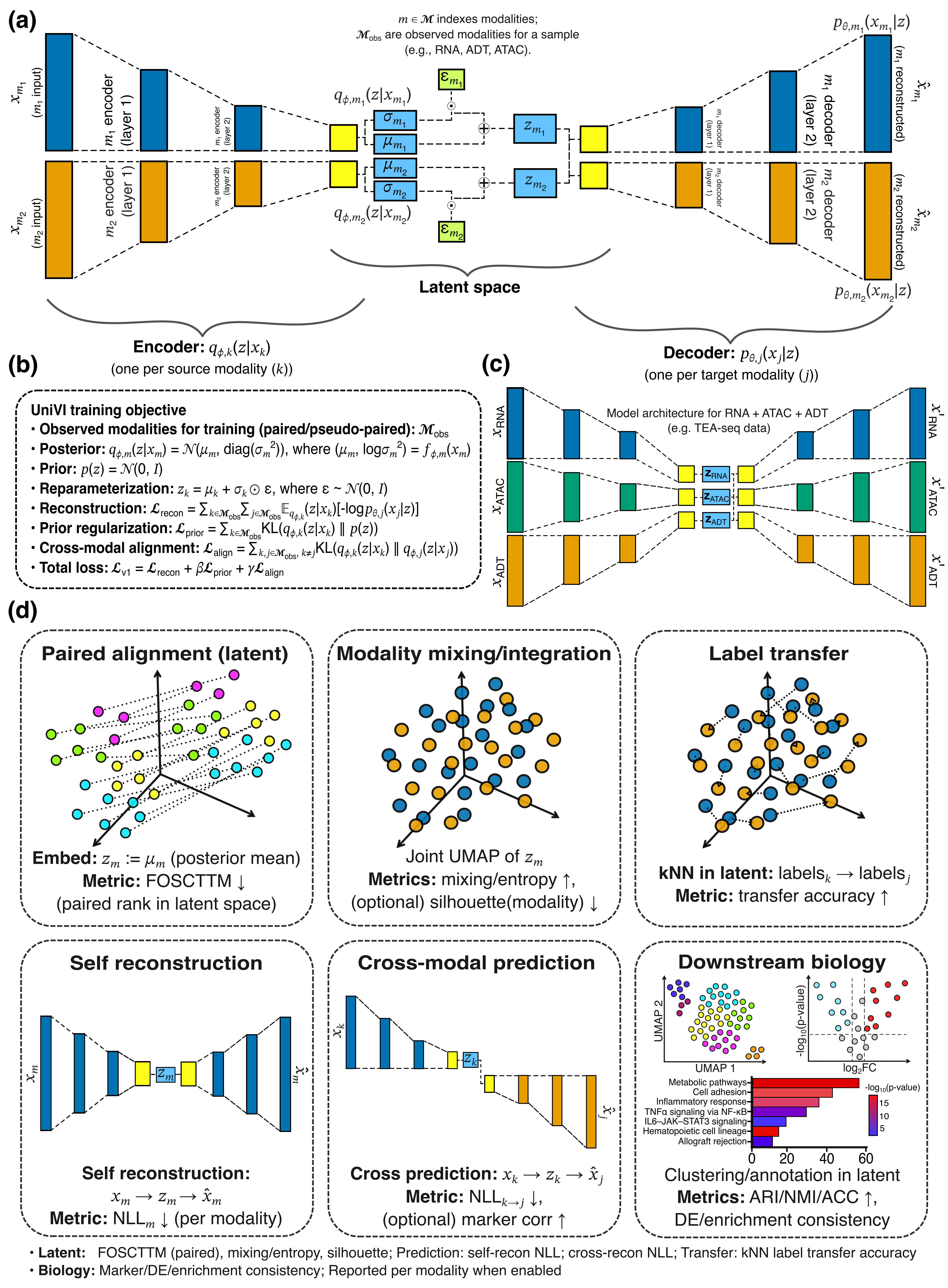
UniVI is a multi-modal variational autoencoder (VAE) toolkit for aligning and integrating single-cell modalities such as RNA, ADT (CITE-seq), ATAC, and coverage-aware / proportion-like assays (e.g., single-cell methylome features).
Common use cases:
- Joint embedding of paired multimodal data (CITE-seq, Multiome, TEA-seq)
- Bridge mapping / projection of unimodal cohorts into a paired latent
- Cross-modal imputation (RNA→ADT, ATAC→RNA, RNA→methylome, …)
- Denoising / reconstruction with likelihood-aware decoders
- Generating biologically-relevant samples due to the generative nature of VAEs
- Evaluation (FOSCTTM, Recall@k, mixing/entropy, label transfer, clustering, basic MoE gating diagnostics)
Advanced/experimental use cases (all optional, model can be run entirely without these):
- Supervised heads (either a decoder classification head or a whole categorical encoder/decoder model VAE, treated as a modality)
- Expanded MoE gating diagnostics (setting a simple gating network during training)
- Transformer encoders (experimental, added for exploratory analysis)
- Fused transformer latent space (even more experimental, added for exploratory analysis/future model expansion)
If you use UniVI in your work, please cite:
Ashford AJ, Enright T, Somers J, Nikolova O, Demir E.
Unifying multimodal single-cell data with a mixture-of-experts β-variational autoencoder framework.
bioRxiv (2025; updated 2026). doi: 10.1101/2025.02.28.640429
@article{Ashford2025UniVI,
title = {Unifying multimodal single-cell data with a mixture-of-experts β-variational autoencoder framework},
author = {Ashford, A. J. and Enright, T. and Somers, J. and Nikolova, O. and Demir, E.},
journal = {bioRxiv},
date = {2025},
doi = {10.1101/2025.02.28.640429},
url = {https://www.biorxiv.org/content/10.1101/2025.02.28.640429},
note = {Preprint (updated 2026)}
}pip install univiUniVI requires PyTorch. If
import torchfails, install PyTorch for your platform/CUDA from PyTorch's official install instructions.
conda install -c conda-forge univi
# or
mamba install -c conda-forge univigit clone https://github.com/Ashford-A/UniVI.git
cd UniVI
conda env create -f envs/univi_env.yml
conda activate univi_env
pip install -e .UniVI expects per-modality AnnData objects.
- Each modality is an
AnnData - For paired settings, modalities share the same cells (
obs_names, same order) - Raw counts often live in
.layers["counts"] - Model inputs typically live in
.X(or.obsm["X_*"]for ATAC LSI) - Model input is a dictionary of these
AnnDataobjects with the dictionary key specifying the modality (e.g.rna,adt,atac). These keys are used later for evaluation functions (cross-reconstruction etc.).
Recommended convention:
.layers["counts"]= raw counts / raw signal.X/.obsm["X_*"]= model input space (log1p RNA, CLR ADT, LSI ATAC, methyl fractions, etc.).layers["denoised_*"]/.layers["imputed_*"]= UniVI outputs
Minimal "notebook path": load paired AnnData → preprocess → train → encode/evaluate → plot.
The sections below walk through a complete CITE-seq (RNA + ADT) example. All patterns generalize to Multiome (RNA + ATAC), TEA-seq (RNA + ADT + ATAC), and any other paired combination supported by UniVI.
import numpy as np
import scanpy as sc
import torch
from torch.utils.data import DataLoader, Subset
from univi import UniVIMultiModalVAE, ModalityConfig, UniVIConfig, TrainingConfig
from univi.data import MultiModalDataset, align_paired_obs_names, collate_multimodal_xy_recon
from univi.trainer import UniVITrainer
collate_multimodal_xy_reconis the required collate function forDataLoaderwhen usingMultiModalDataset. It correctly handles the(x, recon_targets)batch format expected by the trainer, including coverage-aware modalities such as beta-binomial methylome. Always pass it ascollate_fn=collate_multimodal_xy_reconwhen constructing your loaders.
For CITE-seq data:
rna = sc.read_h5ad("path/to/rna_citeseq.h5ad")
adt = sc.read_h5ad("path/to/adt_citeseq.h5ad")For Multiome (RNA + ATAC):
rna = sc.read_h5ad("path/to/rna_multiome.h5ad")
atac = sc.read_h5ad("path/to/atac_multiome.h5ad")For tri-modal TEA-seq / DOGMA-seq / ASAP-seq:
rna = sc.read_h5ad("path/to/rna.h5ad")
adt = sc.read_h5ad("path/to/adt.h5ad")
atac = sc.read_h5ad("path/to/atac.h5ad")After preprocessing, set
.Xto the model input space and keep raw counts in.layers["counts"]. Match thelikelihoodinModalityConfigto your.Xspace (see the likelihood guidance table in step 4).
RNA — log-normalize, select HVGs, scale:
rna.layers["counts"] = rna.X.copy()
rna.var["mt"] = rna.var_names.str.upper().str.startswith("MT-")
sc.pp.calculate_qc_metrics(rna, qc_vars=["mt"], percent_top=None, log1p=False, inplace=True)
sc.pp.normalize_total(rna, target_sum=1e4)
sc.pp.log1p(rna)
rna.raw = rna # snapshot log-space for plotting/DE
sc.pp.highly_variable_genes(rna, flavor="seurat_v3", n_top_genes=2000, subset=True)
sc.pp.scale(rna, max_value=10)ADT — CLR per cell, scale per protein:
adt.layers["counts"] = adt.X.copy()
def clr_per_cell(X):
X = X.toarray() if hasattr(X, "toarray") else np.asarray(X)
logX = np.log1p(X)
return logX - logX.mean(axis=1, keepdims=True)
adt.X = clr_per_cell(adt.layers["counts"])
sc.pp.scale(adt, zero_center=True, max_value=10)ATAC — TF-IDF → LSI, drop first component:
atac.layers["counts"] = atac.X.copy()
def tfidf(X):
X = X.tocsr() if hasattr(X, "tocsr") else X
cell_sum = np.asarray(X.sum(axis=1)).ravel()
cell_sum[cell_sum == 0] = 1.0
tf = X.multiply(1.0 / cell_sum[:, None])
df = np.asarray((X > 0).sum(axis=0)).ravel()
idf = np.log1p(X.shape[0] / (1.0 + df))
return tf.multiply(idf)
X_tfidf = tfidf(atac.layers["counts"])
from sklearn.decomposition import TruncatedSVD
svd = TruncatedSVD(n_components=101, random_state=0)
X_lsi = svd.fit_transform(X_tfidf)
atac.obsm["X_lsi"] = X_lsi[:, 1:] # drop first component (depth correlated)Post-preprocessing: assemble adata_dict
# Sanity check (CITE-seq)
assert rna.n_obs == adt.n_obs and np.all(rna.obs_names == adt.obs_names)
# CITE-seq
adata_dict = {"rna": rna, "adt": adt}
# Multiome
# adata_dict = {"rna": rna, "atac": atac}
# Tri-modal
# adata_dict = {"rna": rna, "adt": adt, "atac": atac}
# Unimodal VAE
# adata_dict = {"rna": rna}
align_paired_obs_names(adata_dict) # ensures matching obs_names and orderAvoiding data leakage: if you want to run UniVI inductively, apply feature selection, scaling, and any learned transforms (e.g., PCA/LSI) on the training set only, then apply the training-set-derived parameters to validation and test sets.
Device detection (CUDA → MPS → XPU → CPU):
device = (
"cuda" if torch.cuda.is_available() else
("mps" if getattr(torch.backends, "mps", None) is not None
and torch.backends.mps.is_available() else
("xpu" if hasattr(torch, "xpu") and torch.xpu.is_available() else
"cpu"))
)Build dataset:
dataset = MultiModalDataset(
adata_dict=adata_dict,
device=None, # dataset yields CPU tensors; model handles GPU transfer
X_key_by_mod={
"rna" : "X", # uses rna.X
"adt" : "X", # uses adt.X
# "atac": "obsm:X_lsi", # uses atac.obsm["X_lsi"]
},
)Train / val / test split (80 / 10 / 10):
n = rna.n_obs
idx = np.arange(n)
rng = np.random.default_rng(0)
rng.shuffle(idx)
n_train = int(0.8 * n)
n_val = int(0.1 * n)
train_idx = idx[:n_train]
val_idx = idx[n_train : n_train + n_val]
test_idx = idx[n_train + n_val :]
# Save split indices for reproducibility
np.savez("splits_seed0.npz", train_idx=train_idx, val_idx=val_idx, test_idx=test_idx)Construct loaders (always pass collate_fn=collate_multimodal_xy_recon):
train_loader = DataLoader(
Subset(dataset, train_idx),
batch_size=256,
shuffle=True,
num_workers=0,
collate_fn=collate_multimodal_xy_recon,
)
val_loader = DataLoader(
Subset(dataset, val_idx),
batch_size=256,
shuffle=False,
num_workers=0,
collate_fn=collate_multimodal_xy_recon,
)
test_loader = DataLoader(
Subset(dataset, test_idx),
batch_size=256,
shuffle=False,
num_workers=0,
collate_fn=collate_multimodal_xy_recon,
)univi_cfg = UniVIConfig(
latent_dim=30,
beta=1.15,
gamma=3.25,
encoder_dropout=0.10,
decoder_dropout=0.05,
encoder_batchnorm=True,
decoder_batchnorm=False,
kl_anneal_start=50,
kl_anneal_end=100,
align_anneal_start=75,
align_anneal_end=125,
modalities=[
ModalityConfig(
name="rna",
input_dim=rna.n_vars,
encoder_hidden=[1024, 512, 256, 128],
decoder_hidden=[128, 256, 512, 1024],
likelihood="gaussian",
),
ModalityConfig(
name="adt",
input_dim=adt.n_vars,
encoder_hidden=[256, 128, 64],
decoder_hidden=[64, 128, 256],
likelihood="gaussian",
),
],
)| Modality | .X / input space |
Recommended likelihood |
|---|---|---|
| RNA | raw counts | "nb" or "zinb" (many zeros) |
| RNA | normalize_total + log1p (+ scale) |
"gaussian" |
| ADT | CLR (+ scale) | "gaussian" |
| ADT | raw counts | "nb" (or "gaussian" post-CLR) |
| ATAC | binarized peaks | "bernoulli" |
| ATAC | raw peak counts | "poisson" |
| ATAC | LSI / reduced | "gaussian" |
| Methylome | fractions / beta values (0–1) | "beta" |
| Methylome | successes + coverage | "beta_binomial" (see methylome quickstart) |
| Methylome | PCA / LSI reduced | "gaussian" |
Manuscript-style tip: Gaussian decoders on normalized feature spaces often produce the most cell-to-cell aligned latent spaces for integration-focused use cases.
Training config and model construction:
train_cfg = TrainingConfig(
n_epochs=3000,
batch_size=256,
lr=1e-3,
weight_decay=1e-4,
device=device,
early_stopping=True,
best_epoch_warmup=50,
patience=50,
log_every=25,
)
model = UniVIMultiModalVAE(
univi_cfg,
loss_mode="v1", # "v1" (manuscript default) or "v2" (lite/fused)
v1_recon="avg",
normalize_v1_terms=True,
).to(device)
loss_modenote:
"v1"(recommended): per-modality posteriors + cross/self/avg reconstruction + posterior alignment. Used in the manuscript."v2"("lite"): fused posterior (MoE/PoE or fused transformer) + per-modality recon + β·KL + γ·L2 alignment. Useful for more loosely-paired/"pseudopaired" settings, but weaker overall cell-to-cell correspondence.
Train:
trainer = UniVITrainer(
model=model,
train_loader=train_loader,
val_loader=val_loader,
train_cfg=train_cfg,
device=device,
)
history = trainer.fit()
print("Best epoch:", getattr(trainer, "best_epoch", None))Save:
ckpt = {
"model_state_dict": model.state_dict(),
"model_config": univi_cfg,
"train_cfg": train_cfg,
"history": getattr(trainer, "history", None),
"best_epoch": getattr(trainer, "best_epoch", None),
}
torch.save(ckpt, "./saved_models/univi_model_state.pt")Load:
import torch
from univi import UniVIMultiModalVAE
ckpt = torch.load("./saved_models/univi_model_state.pt", map_location=device, weights_only=False)
model = UniVIMultiModalVAE(ckpt["model_config"]).to(device)
model.load_state_dict(ckpt["model_state_dict"])
model.eval()
print("Best epoch:", ckpt.get("best_epoch"))UniVI provides utilities to lock down randomness across Python, NumPy, and PyTorch (including CUDA), important for benchmarking and manuscript-level reproducibility.
from univi.utils.seed import set_seed
set_seed(0) # seeds Python, NumPy, torch, and torch.cudaCall set_seed before dataset construction, model initialization, and training to ensure
results are deterministic across runs (given the same hardware and PyTorch version).
You can also snapshot environment + hyperparameter + dataset metadata to a supplemental table:
from univi.diagnostics import export_supplemental_table_s1
export_supplemental_table_s1(
model=model,
train_cfg=train_cfg,
adata_dict=adata_dict,
outpath="./tables/Supplemental_Table_S1.xlsx",
)import numpy as np
import scipy.sparse as sp
import torch
from univi.evaluation import (
encode_adata,
encode_fused_adata_pair,
cross_modal_predict,
denoise_adata,
denoise_from_multimodal,
evaluate_alignment,
reconstruction_metrics,
generate_from_latent,
fit_label_latent_gaussians,
sample_latent_by_label,
evaluate_cross_reconstruction,
)
from univi.plotting import (
set_style,
umap,
umap_by_modality,
compare_raw_vs_denoised_umap_features,
plot_confusion_matrix,
write_gates_to_obs,
plot_moe_gate_summary,
plot_reconstruction_error_summary,
plot_featurewise_reconstruction_scatter,
)
set_style(font_scale=1.2, dpi=150)
def to_dense(X):
return X.toarray() if sp.issparse(X) else np.asarray(X)# Load the saved splits
splits = np.load("splits_seed0.npz")
test_idx = splits["test_idx"]
rna_test = rna[test_idx].copy()
adt_test = adt[test_idx].copy()
assert rna_test.n_obs == adt_test.n_obs
assert np.array_equal(rna_test.obs_names, adt_test.obs_names)Use when you have one observed modality at a time (RNA-only, ADT-only, etc.):
Z_rna = encode_adata(
model,
adata=rna_test,
modality="rna",
device=device,
layer=None, # uses .X by default
X_key="X",
batch_size=1024,
latent="moe_mean", # options: "moe_mean", "moe_sample", "modality_mean", "modality_sample"
random_state=0,
)
rna_test.obsm["X_univi"] = Z_rna
umap(
rna_test,
obsm_key="X_univi",
color=["celltype.l2", "batch"],
legend="outside",
legend_subset_topk=25,
savepath="umap_rna_univi.png",
show=False,
)When you have multiple observed modalities for the same cells:
fused = encode_fused_adata_pair(
model,
adata_by_mod={"rna": rna_test, "adt": adt_test},
device=device,
batch_size=1024,
use_mean=True,
return_gates=True,
return_gate_logits=True,
write_to_adatas=True,
fused_obsm_key="X_univi_fused",
gate_prefix="gate",
)
# fused["Z_fused"] → (n_cells, latent_dim)
# fused["gates"] → (n_cells, n_modalities)
umap(
rna_test,
obsm_key="X_univi_fused",
color=["celltype.l2", "batch"],
legend="outside",
savepath="umap_fused.png",
show=False,
)
umap_by_modality(
{"rna": rna_test, "adt": adt_test},
obsm_key="X_univi_fused",
color=["univi_modality", "celltype.l2"],
legend="outside",
size=8,
savepath="umap_fused_both_modalities.png",
show=False,
)Encode a source modality and decode into a target modality. UniVI automatically handles decoder output types (Gaussian tensor, NB/ZINB/Poisson dicts, Beta/Beta-Binomial dicts) and returns a mean-like prediction matrix ready for evaluation or storage.
# RNA → ADT
adt_hat_from_rna = cross_modal_predict(
model,
adata_src=rna_test,
src_mod="rna",
tgt_mod="adt",
device=device,
layer=None,
X_key="X",
batch_size=512,
use_moe=True,
)
adt_test.layers["imputed_from_rna"] = adt_hat_from_rnaOption A — self-denoise a single modality:
denoise_adata(
model,
adata=rna_test,
modality="rna",
device=device,
out_layer="denoised_self",
overwrite_X=False,
batch_size=512,
)Option B — fused multimodal denoising:
denoise_adata(
model,
adata=rna_test,
modality="rna",
device=device,
out_layer="denoised_fused",
overwrite_X=False,
batch_size=512,
adata_by_mod={"rna": rna_test, "adt": adt_test},
layer_by_mod={"rna": None, "adt": None},
X_key_by_mod={"rna": "X", "adt": "X"},
use_mean=True,
)
compare_raw_vs_denoised_umap_features(
rna_test,
obsm_key="X_univi",
features=["MS4A1", "CD3D", "NKG7"],
raw_layer=None,
denoised_layer="denoised_fused",
savepath="umap_raw_vs_denoised.png",
show=False,
)Basic metrics on two matrices:
true = to_dense(adt_test.X)
pred = adt_test.layers["imputed_from_rna"]
m = reconstruction_metrics(true, pred)
print("MSE mean:", m["mse_mean"])
print("Pearson mean:", m["pearson_mean"])One-call cross-reconstruction evaluation:
rep = evaluate_cross_reconstruction(
model,
adata_src=rna_test,
adata_tgt=adt_test,
src_mod="rna",
tgt_mod="adt",
device=device,
src_layer=None,
tgt_layer=None,
batch_size=512,
feature_names=None, # optionally restrict to a feature subset
)
print(rep["summary"])
plot_reconstruction_error_summary(
rep,
title="RNA → ADT imputation error",
savepath="recon_error_summary.png",
show=False,
)
plot_featurewise_reconstruction_scatter(
rep,
features=["CD3", "CD4", "MS4A1"],
savepath="recon_scatter_selected_features.png",
show=False,
)metrics = evaluate_alignment(
Z1=rna_test.obsm["X_univi"],
Z2=adt_test.obsm["X_univi"],
metric="euclidean",
recall_ks=(1, 5, 10),
k_mixing=20,
k_entropy=30,
labels_source=rna_test.obs["celltype.l2"].to_numpy(),
labels_target=adt_test.obs["celltype.l2"].to_numpy(),
compute_bidirectional_transfer=True,
k_transfer=15,
json_safe=True,
)
plot_confusion_matrix(
np.asarray(metrics["label_transfer_cm"]),
labels=np.asarray(metrics["label_transfer_label_order"]),
title="Label transfer (RNA → ADT)",
normalize="true",
savepath="label_transfer_confusion.png",
show=False,
)UniVI decoders define a per-modality likelihood. Generation works by:
- picking latent samples
z(from the prior or a conditional distribution), - decoding with the target modality's decoder,
- returning a mean-like reconstruction or optionally sampling from the likelihood.
Unconditional generation from standard normal prior:
Xgen = generate_from_latent(
model,
n=5000,
target_mod="rna",
device=device,
z_source="prior",
return_mean=True,
sample_likelihood=False,
)
# Xgen shape: (5000, n_genes)Cell-type–conditioned generation via empirical latent neighborhoods:
Z = rna_test.obsm["X_univi"]
labels = rna_test.obs["celltype.l2"].to_numpy()
# Fit a per-label Gaussian in latent space
label_gauss = fit_label_latent_gaussians(Z, labels)
# Sample latent points for a chosen label
z_B = sample_latent_by_label(label_gauss, label="B cell", n=2000, random_state=0)
# Decode to RNA space
X_B = generate_from_latent(
model,
z=z_B,
target_mod="rna",
device=device,
return_mean=True,
)If you do not have cell-type annotations, you can cluster
Z(e.g., k-means), fit cluster-specific Gaussians, and sample by cluster ID instead.
UniVI can report per-cell modality contribution weights for the analytic fusion path. Two notions of "who contributed" are available:
- Precision-only (always available): derived from each modality's posterior uncertainty.
- Router × precision (when a learnable gating network is present): combines router probabilities with precision to produce contribution weights.
This section applies to analytic fusion (Gaussian experts). If you use the fused transformer posterior, gates may be unavailable or not meaningful.
Compute per-cell contribution weights:
from univi.evaluation import encode_moe_gates_from_tensors
gate = encode_moe_gates_from_tensors(
model,
x_dict={"rna": to_dense(rna_test.X), "adt": to_dense(adt_test.X)},
device=device,
batch_size=1024,
modality_order=["rna", "adt"],
kind="router_x_precision", # falls back to "effective_precision" if no router
return_logits=True,
)
W = gate["weights"] # (n_cells, n_modalities), rows sum to 1
mods = gate["modality_order"]
print("Requested kind:", gate.get("requested_kind"))
print("Effective kind:", gate.get("kind"))
print("Per-modality mean:", gate.get("per_modality_mean"))Write to .obs and plot:
write_gates_to_obs(
rna_test,
gates=W,
modality_names=mods,
gate_prefix="moe_gate",
gate_logits=gate.get("logits"),
)
plot_moe_gate_summary(
rna_test,
gate_prefix="moe_gate",
groupby="celltype.l3",
agg="mean",
savepath="moe_gates_by_celltype.png",
show=False,
)Optionally log gates alongside alignment metrics:
metrics["moe_gates"] = {
"kind": gate.get("kind"),
"requested_kind": gate.get("requested_kind"),
"modality_order": mods,
"per_modality_mean": gate.get("per_modality_mean"),
}For coverage-aware methylome data where you have both methylated counts and total coverage:
rna = sc.read_h5ad("path/to/rna.h5ad")
meth = sc.read_h5ad("path/to/meth.h5ad")
# meth.X → model input (fractions or embeddings)
# meth.layers["meth_successes"] → methylated counts
# meth.layers["meth_total_count"]→ coverage / trials
adata_dict = align_paired_obs_names({"rna": rna, "meth": meth})
recon_targets_spec = {
"meth": {
"successes_layer": "meth_successes",
"total_count_layer": "meth_total_count",
}
}
dataset = MultiModalDataset(
adata_dict=adata_dict,
X_key="X",
device=None,
recon_targets_spec=recon_targets_spec,
)
train_loader = DataLoader(
dataset,
batch_size=256,
shuffle=True,
num_workers=0,
collate_fn=collate_multimodal_xy_recon,
)
univi_cfg = UniVIConfig(
latent_dim=30,
beta=1.15,
gamma=3.25,
modalities=[
ModalityConfig("rna", rna.n_vars, [1024, 512, 256, 128], [128, 256, 512, 1024], likelihood="gaussian"),
ModalityConfig("meth", meth.n_vars, [512, 256, 128], [128, 256, 512], likelihood="beta_binomial"),
],
)
train_cfg = TrainingConfig(n_epochs=2000, batch_size=256, lr=1e-3, weight_decay=1e-4,
device=device, log_every=25)
model = UniVIMultiModalVAE(univi_cfg, loss_mode="v1", v1_recon="avg",
normalize_v1_terms=True).to(device)
trainer = UniVITrainer(model=model, train_loader=train_loader, val_loader=None,
train_cfg=train_cfg, device=device)
history = trainer.fit()When
recon_targetsare present in the batch,UniVITrainerautomatically forwards them intomodel(..., recon_targets=...).
UniVI can project unimodal cells that were not seen during training into the shared latent space ("bridge mapping"), without retraining the model. This is useful for:
- projecting an external RNA-only cohort into a RNA+ADT latent trained on a reference,
- mapping a new patient's ATAC into an existing RNA+ATAC integration,
- any scenario where only a subset of modalities is available at inference time.
# Load a held-out RNA-only cohort (no ADT available)
rna_query = sc.read_h5ad("path/to/rna_query.h5ad")
# Apply the same preprocessing as the reference RNA (HVG subset, scale params, etc.)
# ...
# Encode using only the RNA encoder (modality-specific posterior)
Z_query = encode_adata(
model,
adata=rna_query,
modality="rna",
device=device,
layer=None,
X_key="X",
batch_size=1024,
latent="modality_mean", # use the RNA-specific posterior mean
random_state=0,
)
rna_query.obsm["X_univi"] = Z_queryFine-tuning (optional): If you have a small amount of paired data in the query cohort, you can fine-tune the model on it (typically with a lower learning rate, e.g.,
lr=1e-4, and fewer epochs) to adapt the latent to query-specific batch effects. Seenotebooks/GR_manuscript_reproducibility/UniVI_manuscript_GR-Figure__7__AML_bridge_mapping_and_fine-tuning.ipynbfor an end-to-end example.
For cluster/HPC use or for fully reproducible manuscript runs, UniVI exposes CLI entry points that accept JSON parameter files.
Train from a config JSON:
python scripts/train_univi.py \
--config parameter_files/defaults_cite_seq_scaled_gaussian_v1.json \
--outdir runs/citeseq_v1_run1 \
--data-root /path/to/your/dataEvaluate a saved checkpoint:
python scripts/evaluate_univi.py \
--config parameter_files/defaults_cite_seq_scaled_gaussian_v1.json \
--model-checkpoint runs/citeseq_v1_run1/checkpoints/univi_checkpoint.pt \
--outdir runs/citeseq_v1_run1/evalReproduce all manuscript figures and supplemental tables in one call:
bash scripts/revision_reproduce_all.shExported run directory structure (typical):
runs/<run_name>/
├── checkpoints/
│ ├── univi_checkpoint.pt # model + optimizer + config
│ └── best.pt # best-val checkpoint (if early stopping enabled)
├── eval/
│ ├── metrics.json # machine-readable metrics summary
│ ├── metrics.csv # flat table for comparisons
│ └── plots/ # UMAPs, heatmaps, benchmark figures
├── embeddings/
│ ├── mu_z.npy # fused mean embeddings (cells × latent_dim)
│ ├── modality_mu/ # per-modality embeddings
│ │ ├── rna.npy
│ │ └── adt.npy
│ └── obs_names.txt # row order for safe joins
├── reconstructions/
│ ├── rna_from_rna.npy
│ ├── adt_from_rna.npy # cross-modal imputation
│ └── ...
├── tables/
│ └── Supplemental_Table_S1.xlsx
└── logs/
├── train.log
└── history.csv
You can also invoke the package directly as a module:
python -m univi --helpBy default, UniVI uses MLP encoders. You can switch any modality to a transformer encoder
by setting encoder_type="transformer" and providing a TokenizerConfig and TransformerConfig.
from univi.config import TransformerConfig, TokenizerConfig
univi_cfg = UniVIConfig(
latent_dim=40,
beta=1.0,
gamma=1.25,
modalities=[
ModalityConfig(
name="rna",
input_dim=rna.n_vars,
encoder_hidden=[512, 256, 128], # kept for compatibility; ignored by transformer encoder
decoder_hidden=[128, 256, 512],
likelihood="gaussian",
encoder_type="transformer",
tokenizer=TokenizerConfig(
mode="topk_channels",
n_tokens=512,
channels=("value", "rank", "dropout"),
),
transformer=TransformerConfig(
d_model=256, num_heads=8, num_layers=4,
dim_feedforward=1024, dropout=0.1, attn_dropout=0.1,
activation="gelu", pooling="mean",
),
),
ModalityConfig(
name="adt",
input_dim=adt.n_vars,
encoder_hidden=[128, 64],
decoder_hidden=[64, 128],
likelihood="gaussian",
encoder_type="mlp",
tokenizer=TokenizerConfig(mode="topk_scalar", n_tokens=min(32, adt.n_vars)),
),
],
)For ATAC tokenizers, you can optionally incorporate genomic coordinate embeddings and distance-aware attention bias to encourage local regulatory context:
# Tokenizer config with coordinate embeddings enabled
TokenizerConfig(
mode="topk_channels",
n_tokens=512,
channels=("value", "rank", "dropout"),
use_coord_embedding=True,
n_chroms=<num_chromosomes>,
coord_scale=1e-6,
)
# Provide peak coordinates to the trainer
feature_coords = {
"atac": {
"chrom_ids": chrom_ids_long, # (F,) integer chromosome IDs
"start": start_bp, # (F,) start positions in bp
"end": end_bp, # (F,) end positions in bp
}
}
attn_bias_cfg = {
"atac": {
"type": "distance",
"lengthscale_bp": 50_000.0,
"same_chrom_only": True,
}
}
trainer = UniVITrainer(
model,
train_loader,
val_loader=val_loader,
train_cfg=TrainingConfig(...),
device=device,
feature_coords=feature_coords,
attn_bias_cfg=attn_bias_cfg,
)
trainer.fit()Instead of analytic MoE/PoE fusion of per-modality posteriors, you can use a single
transformer that sees concatenated tokens from all modalities and outputs a single
fused posterior q(z | all modalities):
from univi.config import TransformerConfig
univi_cfg = UniVIConfig(
latent_dim=40,
beta=1.0,
gamma=1.25,
modalities=[...],
fused_encoder_type="multimodal_transformer",
fused_modalities=("rna", "adt", "atac"),
fused_require_all_modalities=True, # fall back to MoE if any modality is missing
fused_transformer=TransformerConfig(
d_model=256, num_heads=8, num_layers=4,
dim_feedforward=1024, dropout=0.1, attn_dropout=0.1,
activation="gelu", pooling="cls",
),
)Encoding with the fused transformer posterior:
mu, logvar, z = model.encode_fused(
{"rna": X_rna, "adt": X_adt, "atac": X_atac},
use_mean=True,
)When
fused_require_all_modalities=True, the model falls back to MoE fusion automatically for cells missing one or more modalities.
By default, fusion uses pure precision weighting. You can add a learnable gate network that produces data-driven per-cell modality trust scores:
univi_cfg = UniVIConfig(
...,
use_moe_gating=True,
moe_gating_type="per_modality", # or "shared"
moe_gating_hidden=[128, 64],
moe_gating_dropout=0.1,
moe_gating_batchnorm=True,
moe_gate_eps=1e-6,
)Retrieve gates after encoding:
mu, logvar, z, gates, gate_logits = model.encode_fused(
x_dict,
use_mean=True,
return_gates=True,
return_gate_logits=True,
)
# gates: (n_cells, n_modalities)UniVI supports any number of supervised heads trained jointly with the VAE objective. Heads can be categorical, binary, or adversarial (domain confusion via gradient reversal).
from univi.config import ClassHeadConfig
univi_cfg = UniVIConfig(
...,
class_heads=[
ClassHeadConfig(
name="celltype",
n_classes=n_celltypes,
loss_weight=1.0,
ignore_index=-1,
from_mu=True,
warmup=0,
),
ClassHeadConfig(
name="batch",
n_classes=n_batches,
loss_weight=0.2,
ignore_index=-1,
from_mu=True,
warmup=10,
adversarial=True,
adv_lambda=1.0,
),
ClassHeadConfig(
name="TP53_mut",
n_classes=2,
loss_weight=0.5,
ignore_index=-1,
from_mu=True,
),
],
)
model.set_head_label_names("celltype", list(rna.obs["celltype"].astype("category").cat.categories))Pass labels as a dict during training:
y = {
"celltype": torch.tensor(celltype_codes[batch_idx], device=device),
"batch": torch.tensor(batch_codes[batch_idx], device=device),
"TP53_mut": torch.tensor(tp53_flags[batch_idx], device=device),
}
out = model(x_dict, epoch=epoch, y=y)
loss = out["loss"]
loss.backward()Predict heads after training:
model.eval()
with torch.no_grad():
probs = model.predict_heads(x_dict, return_probs=True)
for head_name, P in probs.items():
print(head_name, P.shape)
# Inspect head config metadata
meta = model.get_classification_meta()
print(meta)Rather than a prediction head, you can treat a categorical variable as a generative modality
with its own encoder q(z|y) and decoder p(y|z), participating fully in fusion and
posterior alignment.
univi_cfg = UniVIConfig(
...,
modalities=[
ModalityConfig("rna", rna.n_vars, [512, 256, 128], [128, 256, 512], likelihood="nb"),
ModalityConfig("adt", adt.n_vars, [128, 64], [64, 128], likelihood="nb"),
ModalityConfig(
name="celltype_mod",
input_dim=n_celltypes, # number of classes
encoder_hidden=[128, 64],
decoder_hidden=[64, 128],
likelihood="categorical",
),
],
)Pass integer labels (or one-hot) via x_dict["celltype_mod"] during training.
Unlabeled cells should use -1 (or your configured ignore_index). This is distinct
from classification heads: the categorical modality is part of the generative model, not
an auxiliary output.
Semi-supervised alignment where label information is fused as an additional Gaussian expert:
univi_cfg = UniVIConfig(
...,
use_label_encoder=True,
n_label_classes=n_celltypes,
label_encoder_warmup=50, # epochs before label injection starts
label_moe_weight=1.0,
unlabeled_logvar=10.0, # large → label contributes little when missing
)
# At encode time:
mu, logvar, z = model.encode_fused(
x_dict,
epoch=current_epoch,
y={"label": y_ids},
inject_label_expert=True,
)When modalities have very different feature dimensionalities (RNA: 2k–20k; ADT: 30–200; ATAC-LSI: ~50–500), reconstruction losses summed over features can cause high-dimensional modalities to dominate gradients. UniVI supports per-modality normalization:
model = UniVIMultiModalVAE(
univi_cfg,
loss_mode="v1",
v1_recon="avg",
normalize_v1_terms=True,
recon_normalize_by_dim=True, # divide recon loss by D^power
recon_dim_power=0.5, # 0.5 = soft correction; 1.0 = full per-feature mean
).to(device)Alternatively, set per-modality weights directly in ModalityConfig:
ModalityConfig(
name="rna",
...,
recon_weight=1.0,
)
ModalityConfig(
name="adt",
...,
recon_weight=2.0, # upweight ADT relative to RNA
)UniVI ships with modality-specific hyperparameter search scripts:
from univi.hyperparam_optimization import (
run_citeseq_hparam_search,
run_multiome_hparam_search,
run_teaseq_hparam_search,
run_rna_hparam_search,
run_atac_hparam_search,
run_adt_hparam_search,
)Or via the CLI:
python scripts/run_multiome_hparam_search.py \
--config parameter_files/defaults_multiome_v1.json \
--outdir runs/hparam_search_multiome \
--n-trials 50See univi/hyperparam_optimization/ and the grid-sweep notebooks in
notebooks/GR_manuscript_reproducibility/ for end-to-end examples with result aggregation.
UniVI/
├── README.md # Project overview, installation, quickstart
├── LICENSE # MIT license text file
├── pyproject.toml # Python packaging config (pip / PyPI)
├── assets/ # Static assets used by README/docs
│ └── figures/ # Schematic figure(s) for repository front page
├── conda.recipe/ # Conda build recipe (for conda-build)
│ └── meta.yaml
├── envs/ # Conda/mamba environment examples you can use
│ ├── UniVI_working_environment.yml
│ ├── UniVI_working_environment_v2_full.yml
│ ├── UniVI_working_environment_v2_minimal.yml
│ ├── univi_env.yml
│ └── univi_v0.4.7.yml # Most recent working environment (v0.4.7) - recommended
├── data/ # Small example data notes (datasets are typically external)
│ └── README.md # Notes on data sources / formats
├── notebooks/ # End-to-end Jupyter Notebook analyses and examples
│ ├── GR_manuscript_reproducibility/ # Reproduce figures from revised manuscript (in progress for Genome Research; bioRxiv manuscript v2)
│ │ ├── UniVI_manuscript_GR-Figure__2__CITE_paired.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__3__CITE_paired_biological_latent.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__4__Multiome_paired.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__5__Multiome_bridge_mapping_and_fine-tuning.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__6__TEA-seq_tri-modal.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__7__AML_bridge_mapping_and_fine-tuning.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__8__benchmarking_against_pytorch_tools.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__8__benchmarking_against_R_tools.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__8__benchmarking_merging_and_plotting_runs.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__9__paired_data_ablation_and_computational_scaling_performance.ipynb
│ │ ├── UniVI_manuscript_GR-Figure__9__paired_data_ablation_and_computational_scaling_performance_compile_plots_from_results_df.ipynb
│ │ ├── UniVI_manuscript_GR-Figure_10__cell_population_ablation_MoE.ipynb
│ │ ├── UniVI_manuscript_GR-Figure_10__cell_population_ablation_MoE_compile_plots_from_results_df.ipynb
│ │ ├── UniVI_manuscript_GR-Supple_____grid-sweep.ipynb
│ │ └── UniVI_manuscript_GR-Supple_____grid-sweep_compile_plots_from_results_df.ipynb
│ └── UniVI_additional_examples/ # Additional examples of UniVI workflow functionality
│ ├── Multiome_NB-RNA-counts_Poisson_or_Bernoulli-ATAC_peak-counts_Peak_perturbation_to_RNA_expression_cross-generation_experiment.ipynb
│ └── scNMT-seq_mouse_gastrulation_feature-level_integration_example.ipynb
├── parameter_files/ # JSON configs for model + training + data selectors
│ ├── defaults_*.json # Default configs (per experiment)
│ └── params_*.json # Example “named” configs (RNA, ADT, ATAC, methylome, etc.)
├── scripts/ # Reproducible entry points (revision-friendly)
│ ├── train_univi.py # Train UniVI from a parameter JSON
│ ├── evaluate_univi.py # Evaluate trained models (FOSCTTM, label transfer, etc.)
│ ├── benchmark_univi_citeseq.py # CITE-seq-specific benchmarking script
│ ├── run_multiome_hparam_search.py
│ ├── run_frequency_robustness.py # Composition/frequency mismatch robustness
│ ├── run_do_not_integrate_detection.py # “Do-not-integrate” unmatched population demo
│ ├── run_benchmarks.py # Unified wrapper (includes optional Harmony baseline)
│ └── revision_reproduce_all.sh # One-click: reproduces figures + supplemental tables
└── univi/ # UniVI Python package (importable as `import univi`)
├── __init__.py # Package exports and __version__
├── __main__.py # Enables: `python -m univi ...`
├── cli.py # Minimal CLI (e.g., export-s1, encode)
├── pipeline.py # Config-driven model+data loading; latent encoding helpers
├── diagnostics.py # Exports Supplemental_Table_S1.xlsx (env + hparams + dataset stats)
├── config.py # Config dataclasses (UniVIConfig, ModalityConfig, TrainingConfig)
├── data.py # Dataset wrappers + matrix selectors (layer/X_key, obsm support)
├── evaluation.py # Metrics (FOSCTTM, mixing, label transfer, feature recovery)
├── matching.py # Modality matching / alignment helpers
├── objectives.py # Losses (ELBO variants, KL/alignment annealing, etc.)
├── plotting.py # Plotting helpers + consistent style defaults
├── trainer.py # UniVITrainer: training loop, logging, checkpointing
├── interpretability.py # Helper scripts for transformer token weight interpretability
├── figures/ # Package-internal figure assets (placeholder)
│ └── .gitkeep
├── models/ # VAE architectures + building blocks
│ ├── __init__.py
│ ├── mlp.py # Shared MLP building blocks
│ ├── encoders.py # Modality encoders (MLP + transformer + fused transformer)
│ ├── decoders.py # Likelihood-specific decoders (NB, ZINB, Gaussian, Beta, Binomial/Beta-Binomial, etc.)
│ ├── transformer.py # Transformer blocks + encoder (+ optional attn bias support)
│ ├── tokenizer.py # Tokenization configs/helpers (top-k / patch)
│ └── univi.py # Core UniVI multi-modal VAE
├── hyperparam_optimization/ # Hyperparameter search scripts
│ ├── __init__.py
│ ├── common.py
│ ├── run_adt_hparam_search.py
│ ├── run_atac_hparam_search.py
│ ├── run_citeseq_hparam_search.py
│ ├── run_multiome_hparam_search.py
│ ├── run_rna_hparam_search.py
│ └── run_teaseq_hparam_search.py
└── utils/ # General utilities
├── __init__.py
├── io.py # I/O helpers (AnnData, configs, checkpoints)
├── logging.py # Logging configuration / progress reporting
├── seed.py # Reproducibility helpers (seeding RNGs)
├── stats.py # Small statistical helpers / transforms
└── torch_utils.py # PyTorch utilities (device, tensor helpers)
MIT License — see LICENSE.
-
Questions / comments: open a GitHub Issue with the
questionlabel (or use Discussions) -
Bug reports: include:
- UniVI version:
python -c "import univi; print(univi.__version__)" - minimal notebook/code snippet
- stack trace + OS/CUDA/PyTorch versions
- UniVI version:




